Improving the automation of our Machine Learning pipelines
Our Chief Architect, Olly, talks us through a gap we found when improving our Pachyderm integration in the Adarga Data Science Workbench, and the open-source code we created to plug it; all part of our infrastructure that enables high-velocity Data Science and the creation of our AI software.
Here at Adarga, we make extensive use of Pachyderm within our R&D environment. Our team of data scientists uses it to experiment with new models and flows, as well as running the usual batch of training jobs, etc. Pachyderm gives us confidence that we are keeping careful track of the data we use and its provenance, as well as giving us a convenient way of expressing the pipeline of steps we wish to apply to it. It is at the heart of what we refer to as our Data Science Workbench.
Despite data scientists having free rein to run whatever experiments take their fancy in the Data Science Workbench, we like to impose the same engineering discipline on it that we have on all our other Kubernetes clusters, and to that end, we make extensive use of ArgoCD to power a GitOps approach to cluster management. This is a key part of our internal MLOps story; not only does it mean we have a good understanding - and audit trail - of what is running in the Data Science Workbench, but it also reduces the burden - cognitive load - on our data scientists in terms of launching new services by reducing their need to understand Kubernetes under the hood, and it allows us to provide automated paths from the Data Science Workbench to our production environments.
By providing them with abstractions such as SeldonDeployment objects, we reduce their interactions down to stewardship of a small number of easy-to-understand declarative manifests. But this caused an issue when we first integrated Pachyderm. Despite it having a nice declarative syntax for defining pipelines, the only method provided for interacting with them was via the pachctl command line. We decided we needed a way to get the Pachyderm pipeline definitions into Kubernetes so that we could manage their lifecycle from within the cluster, triggered by ArgoCD. After a few iterations that were somewhat fragile built using various combinations of config maps and jobs, we decided to bite the bullet and create a controller that handles this. And I’m pleased to say it’s now working very well for us and we have open-sourced the code.
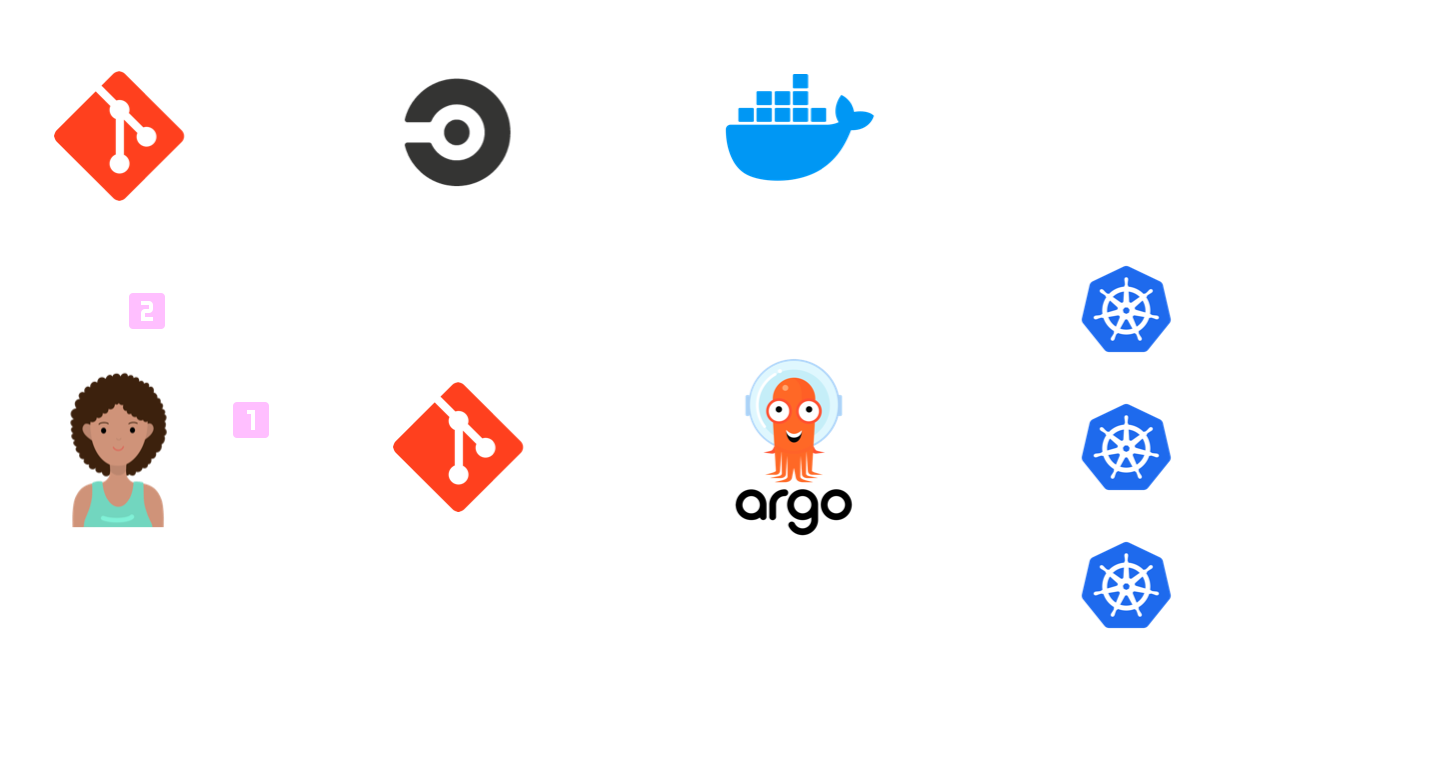
How the controller works
The key difference from a normal Pachyderm flow is that we have introduced a custom PachydermPipeline object. This holds the same information as a normal pachyderm pipeline definition file but is held in Kubernetes. Our controller monitors these objects and will reflect any changes into Pachyderm itself using the pachd API.
On the surface, this doesn’t appear to add much value. The process for managing Pachyderm pipelines is largely the same; the only tangible difference is what format you store them in and what command you run to enact the change:

However, this subtle difference allows us to leverage our existing ArgoCD tooling to apply GitOps principles to the pipelines. ArgoCD is designed to spot a change to a Kubernetes manifest in a git repository and apply that change to the cluster. By switching the pipeline definitions to be Kubernetes manifests, it can apply its logic to those too.
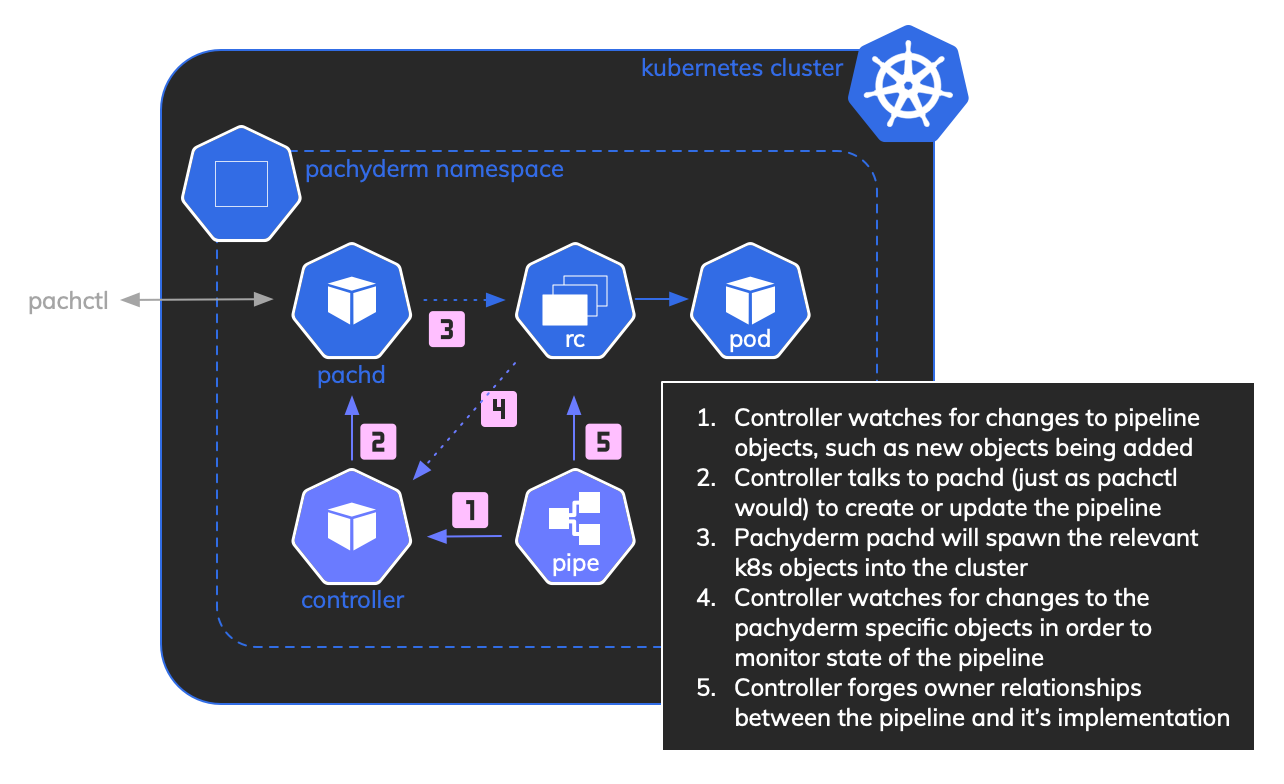
Having pipelines as first-class objects in Kubernetes gives you an additional angle for querying against (although pachctl knows way more about the pipelines than the controller does, so should still be the first port of call):
1 % kubectl get pipe 2 NAME AGE CREATED CONDITION 3 base-entity-recogniser 11d Success Stopped 4 doc2vec-generator 11d Success Running 5 zero-shot-classifier 11d CreationError Missing
and the controller makes the effort to connect these new objects correctly with the Pachyderm-generated objects of the pipeline for better visibility, either on the command line:
1 % kubectl tree pipe doc2vec-generator 2 NAMESPACE NAME READY REASON AGE 3 pach-feed PachydermPipeline/doc2vec-generator - 11d 4 pach-feed └─ReplicationController/pipeline-doc2vec-generator-v5 - 9h 5 pach-feed └─Pod/pipeline-doc2vec-generator-v5-rj5cp TRUE 9h
or in the ArgoCD visualization of the cluster:

Current status
At the time of writing, the controller has sufficient functionality for our internal needs and is production deployed into our internal Data Science Workbench. It doesn’t support the full Pachyderm pipeline spec - just the subset that we use, so that’s an area we hope to find time to improve (PRs welcome!). Our next plan in terms of functionality, however, is to add support for multiple Pachyderm clusters within a single Kubernetes cluster and to integrate the Argo project’s new notification engine so we can deliver Slack notifications to the users when pipeline lifecycle events occur.